Named after chemist Rosalind Franklin, whose work contributed to understanding the structure of DNA, GPT-Rosalind is focused on tasks in biology, drug discovery, and translational medicine. According to OpenAI, it can support researchers with evidence synthesis, hypothesis generation, experimental design, and other multi-step scientific workflows.
Unlike general-purpose GPT models, GPT-Rosalind is positioned as a system optimized for scientific research workflows. OpenAI says it is better at reasoning about molecules, proteins, genes, signaling pathways, and disease-related biology, while also using scientific tools and databases more effectively in multi-step processes. Supported use cases include literature review, sequence-function interpretation, experiment planning, and data analysis.
Internal benchmarks show gains, but independent validation is still needed
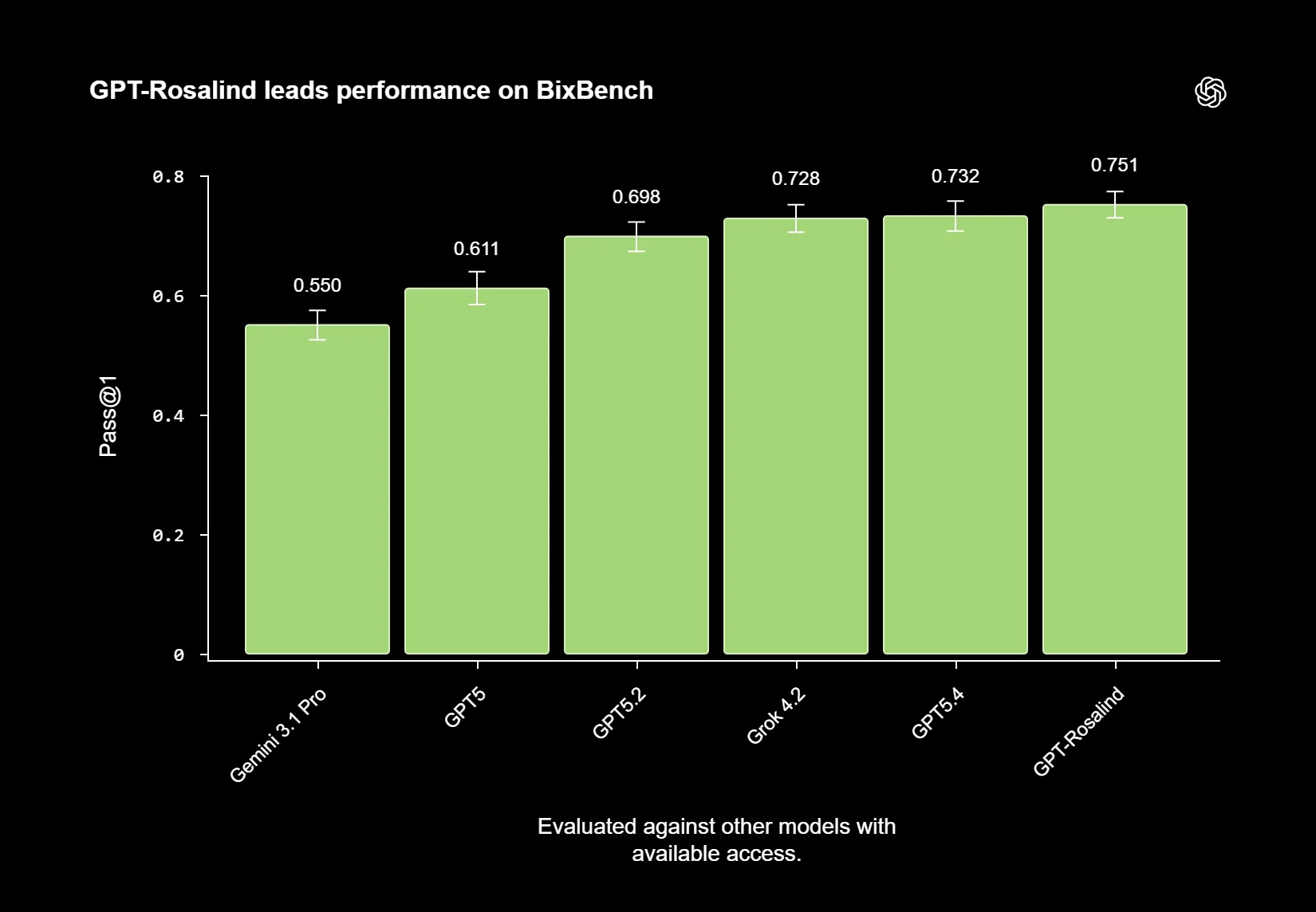
In OpenAI’s internal evaluations, GPT-Rosalind reportedly outperforms earlier models including GPT-5, GPT-5.2, and GPT-5.4 across five tested categories: chemistry, biochemistry and protein understanding, phylogenetics, experiment design and analysis, and tool use. The largest improvements were reported in chemistry and experimental design.
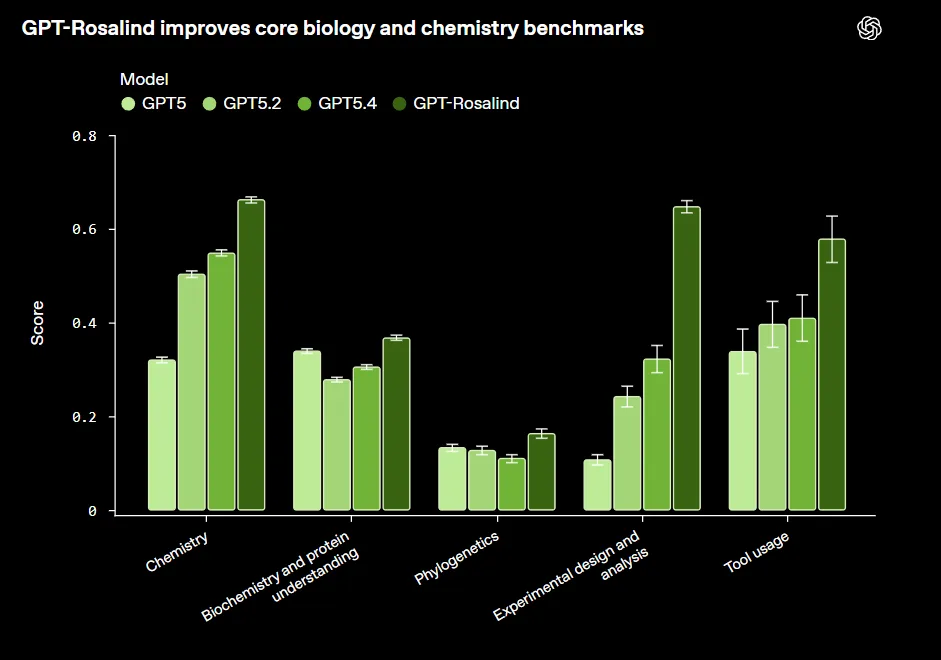
On the public BixBench benchmark for bioinformatics and data analysis, GPT-Rosalind achieved a Pass@1 score of 0.751. OpenAI says this places it ahead of GPT-5.4 (0.732), GPT-5 (0.728), Grok 4.2 (0.698), and Gemini 3.1 Pro (0.550).
On LABBench2, which includes tasks such as literature search, database access, sequence manipulation, and protocol design, GPT-Rosalind reportedly outperformed GPT-5.4 in 6 of 11 tasks. OpenAI says the strongest improvement appeared in CloningQA, a task requiring full design of DNA and enzyme reagents for molecular cloning protocols.
Open-source plugin connects more than 50 scientific data sources
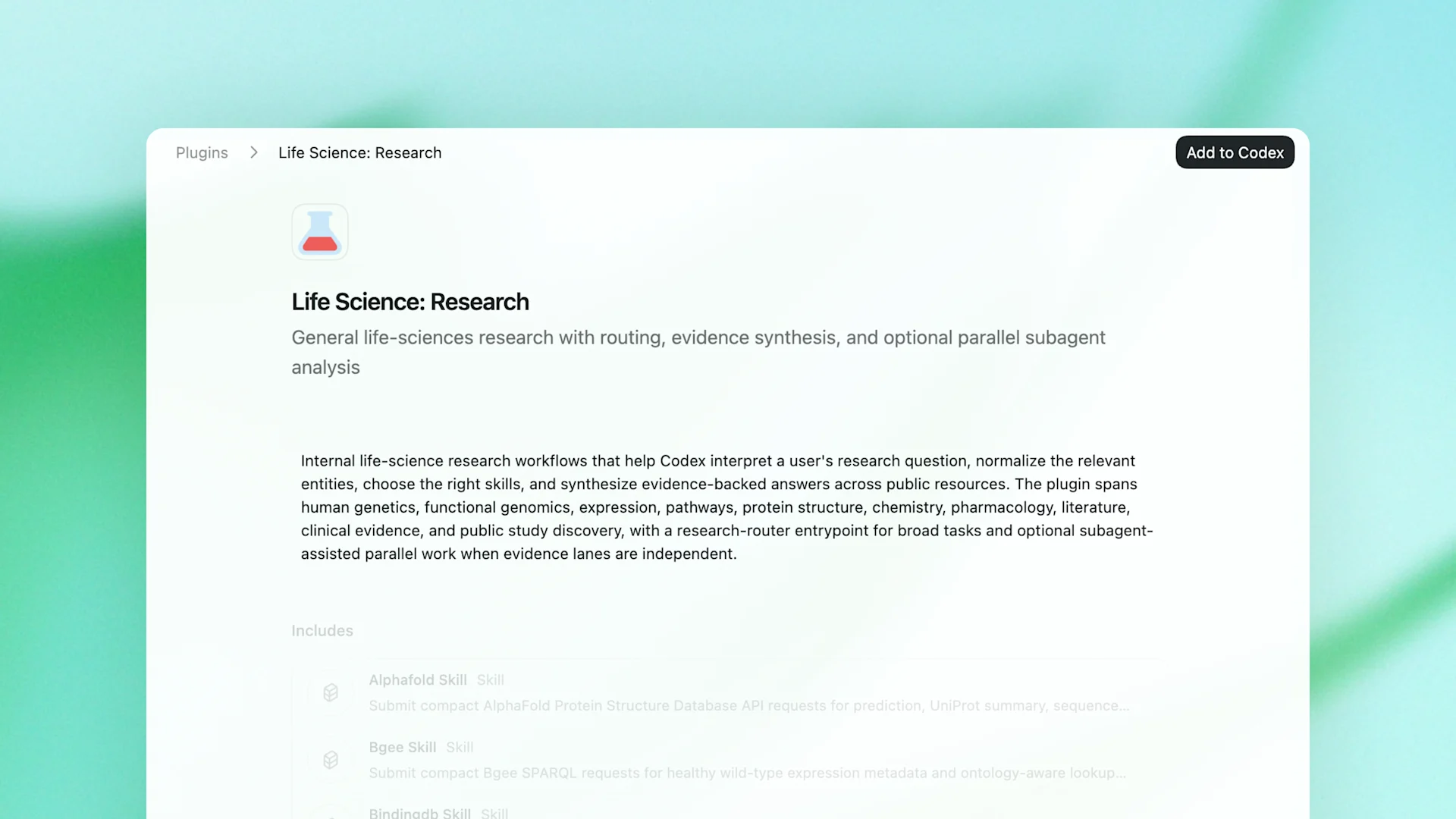
Alongside the model, OpenAI has released a free Life Sciences Research plugin for Codex on GitHub. The plugin provides modular skills for common research workflows and connects models to more than 50 public multi-omics databases, literature sources, and biology tools.
The supported sources include areas such as human genetics, functional genomics, protein structure, biochemistry, clinical evidence, and public study discovery. According to OpenAI, the plugin acts as an orchestration layer for broad, ambiguous, and multi-step scientific questions. Enterprise users can pair it with GPT-Rosalind, while other users can access it with OpenAI’s standard models.
Limited research preview and Trusted Access Program
GPT-Rosalind is available as a research preview in ChatGPT, Codex, and the API, but only for qualified Enterprise customers in the United States through a Trusted Access Program. During the preview phase, OpenAI says usage will not consume existing credits or tokens. Pricing and wider availability have not yet been announced.
Access is based on three conditions:
legitimate scientific research with clear public benefit, strong governance and misuse-prevention controls, and restricted access for approved users in secure managed environments.
First model in a planned life sciences series
OpenAI describes GPT-Rosalind as the first release in a planned series of life sciences models. The company says it intends to keep improving biochemical reasoning for tool-intensive and long-horizon research workflows.
According to OpenAI, early users and partners include Amgen, Novo Nordisk, Moderna, Thermo Fisher Scientific, Oracle Health and Life Sciences, NVIDIA, the Allen Institute, Benchling, and the UCSF School of Pharmacy.


 ES
ES  EN
EN